When analyzing genomic data, we first need to align to the genome. There are a lot of possible choices in this, including BWA (medium choice), stampy (very accurate) and bowtie2 (very fast). Recently a new aligner came out, NextGenMap. It claims to be both faster and deal with divergent read data better than other methods. Continue reading
Tag Archives: snp calling
The limits of GBS sample size
Image
I’ve been doing work on a stickleback GBS dataset and we’re trying to figure out how many samples we can cram into a single lane of illumina. I did some analyses which people may find useful. It’s unclear how applicable the recommendations are for sunflower which seems to have more problems than fish.
Take home message, with 25% less data you lose around 15% of the genotype calls, but up to 50% of the SNPs if you use a stringent coverage filter, due to how the lost data is distributed among loci and individuals.
SNP calling with GATK warning
I’ve come to the realization that I’ve been making a mistake in my SNP calling using GATK. I’ve been filtering for site quality not for individual genotype quality. This is a critical difference and leads to some dubious SNPs being called.
It works like this. For each position in the genome, GATK will use all the data together to call whether an alternate allele exists. This is represented by the phred scaled QUAL score. A high QUAL score means that the alternate allele exists in in your dataset. This number is often very high if you’re analyzing a whole plate of GBS because it takes into account all the data from all the individuals. I’ve been filtering based on this number.
There is also a MQ score, which is the mapping quality for all the reads at that site.
For each individual, if there are any reads at that site GATK will output a genotype call (i.e. 0/0, 0/1, 1/1). It also outputs the number of reads supporting each allele, the QT score (which is the phred scaled probability of that the genotype call being correct) and the likelihoods of each possible genotype. When you use select variants in GATK, it filters sites but not genotypes (so you can have a site that is good, but individual genotype calls at that site that are bad).
If you use vcf-to-tab to convert your vcf to tab separated it will accept every genotype call. This means that for some individuals you will have 1 read supporting the reference and it will be called as 0/0. The QT score will be 3, which is incredibly low, but vcf-to-tab ignores that information and calls it. Therefore a proportion of your SNPs are highly questionable. This likely contributes to the heterozygote loss seen in GBS data.
Sam and Chris wrote a script that converts a vcf file to tab but also makes sure that the quality (QUAL, QT, MQ, and depth) is good. I’ll post it if they’re OK with it be up.
Edit: Here are the scripts. They are identical except one outputs in IUPAC format (AT=W, AC=M, etc) and the other outputs in the standard double nucleotide format (AA, AT, etc).vcf2vertical_dep_GATK-UG vcf2vertical_dep_GATK
Also note that sometimes GATK will output a genotype call that looks like:
./.:.:3
While normally they should look like:
./. or 0/0:17,0:18:45:0,45,353
This is saying that there are three reads at this position, but the quality is so bad that GATK is not calling the genotype. If you have these, the scripts will throw an error but will correctly output an NN for that genotype. So if you get a normal output and many errors, it’s probably OK.
Where does all the GBS data go?
Why do we get seemingly few SNPs with GBS data?
Methods: Used bash to count the occurrences of tags. Check your data and let me know what you find:
cat Demutliplexed.fastq | sed -n '2~4p' | cut -c4-70 | sort | uniq -c | grep -v N | sed 's/\s\s*/ /g' > TagCounts
Terminology: I have tried to use tags to refer to a particular sequence and reads as occurrences of those sequences. So a tag may be found 20 times in a sample (20 reads of that sequence).
Findings: Tag repeat distribution
Probably the key thing and a big issue with GBS is the number of times each tag gets sequence. Most sites get sequenced once but most tags get sequence many many times. It is hard to see but here are histograms of the number of times each tags occur for a random sample (my pet11): All tags histogram (note how long the x axis is), 50 tags or less Histogram. Most tags occur once – that is the spike at 1. The tail is long and important. Although only one tag occurs 1088 times it ‘takes up’ 1088 reads.
How does this add up?
In this sample there are 3,453,575 reads. These reads correspond to 376,036 different tag sequences. This would mean (ideally) ~10x depth of each of the tags. This is not the case. There are a mere 718 tags which occur 1000 or more times but they account for 1394905 reads. That is 40% of the reads going to just over 700 (potential) sites. I’ve not looked summarized more samples using the same method but I am sure it would to yield the same result.
Here is an example: Random Deep Tag. Looking at this you can see that the problem is worse than just re-sequencing the same tag many times but you introduce a large number of tags that are sequencing and/or PCR errors that occur once or twice (I cut off many more occurances here).
Conclusion: Poor/incomplete digestion -> few sites get adapters -> they get amplified like crazy and then sequenced.
Update 1:
Of the > 3.4Million tags that are in the set of 18 samples I am playing with only 8123 are in 10 or more of the samples.
For those sites with >10 scored the number of times a tag is sequenced is correlated between samples. The same tags are being sequenced repeatedly in the different samples.
Update 2:
As requested the ‘connectivity’ between tags. This is the number of 1bp miss matches each tag has. To be included in this analysis a tag must occur at least 5 times in 3 individuals. Here is the figure. So most tags have zero matches a smaller number have one and so on. This actually looks pretty good – at least how I am thinking about it now. This could mean that the filtering criteria works. If errors were still being included I would expect tags with one match (the actual sequence) to occur more than ones with zero.
GBS, coverage and heterozygosity. Part 3
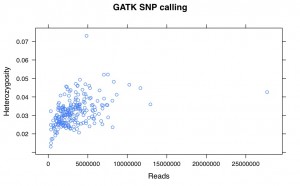
I’ve recalled the SNPs for my Helianthus exilis population genetics for a third time. This time I’m using GATK. This is aligned using BWA to the Nov22k22 reference.
This is two plates of GBS (96-plex each) plus 34 H. annuus samples from G. Baute (also 96-plex GBS). Three exilis samples were removed because they had little or no reads. Reads were trimmed for adapters and quality using trimmomatic (and the number of reads kept after trimming are used here).
So, there is a relationship between number of reads and resulting heterozygosity. It makes some sense because you need a higher number of reads to call a heterozygote than a homozygote. It’s not as bad as what was happening when we were calling snps using the maximum likelihood script. Using a linear model, number of reads accounts for %16 of the variation in heterozygosity.
If you compare this to my previous posts, the heterozygosity is vastly lower for all samples. That is because I previously looked at variable sites with a certain amount of coverage, and here are all sites. So a majority of these sites are invariant.
SNP calling VI – Variant filtering
Filtering the variants is a critical step in the pipeline, as most of the SNP callers are very inclusive by default. Below are the criteria I have used to filter SNPs:
SNP calling V – SNP and genotype calling
The next step in the pipeline after the alignments are conducted and adjusted is to call the SNPs and genotypes. There are many programs that will do this such as the ML script that many people in the Rieseberg lab use, freeBayes, mpileup and bcftools, as well as the GATK UnifiedGenotyper. To bench mark the alignment steps in these SNP calling blog posts I used mpileup and bcftools. For example:
SNP calling IV – Base quality score recalibration
The best practices protocol from Broad highly recommends base quality recalibration (http://gatkforums.broadinstitute.org/discussion/44/base-quality-score-recalibration-bqsr). Apparently, the base quality scores off the machine are not very accurate and can provide false confidence in the base calls and consequently influence SNP calling. This tool will adjust average quality scores and also adjust the scores depending on the machine cycle and sequence context (i.e. start versus end of read, type of dinucleotide repeat). After recalibration, the quality score is closer to its actual probability of mismatching the reference genome.
To run this you need a database of known SNPs so the tool can assess the error rate at other sites based on the actual sequence data. Unlike humans, which this tool was designed for, most species do not have comprehensive SNP databases. However, such a table can be created by identifying SNPs in a preliminary run without the base quality score recalibration. I included any SNP that had a Qual score of 20 in my SNP vcf file.
You can see below that it had a large effect on the number of SNPs that I called. With the BAQ filter off 263,360 SNPS overlapped between the methods. 14,591 were unique to the recalibrated alignments and 131,354 were unique to the alignments without base quality recalibration.
SNP calling III – The indel problem
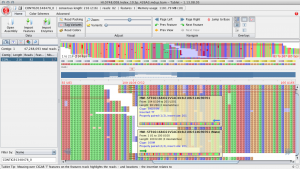
SNPs can often be falsely called around indels. In particular, if indels are near the start or end of a read the read is often incorrectly aligned. Here is an example of how things can go wrong from a spruce alignment. Below, at site 109 there is a TT insertion in the reads relative to the reference and there is no true SNP at this site. However, this individual was called as a het (C/T) because of the reads that end in the middle of the insertion. For these reads the penalty for a single mismatch to the reference is less than introducing a gap.
SNP calling II – Creating a reference for GATK and Picard
Using GATK and Picard was difficult because we had to keep modifying our reference (and therefore redo the alignments each time) to use these programs efficiently. Below I list the things we had to do to the reference to get them to work:
SNP calling I – alignment programs and PCR duplicates
I have tested various SNP calling methods using exome re-sequencing data from 12 interior spruce samples. I tried Bowtie2, BWA (mem), Picard (mark duplicates) and GATK for indel realignment and base quality recalibration. For SNP calling I used mpileup with and without BAQ as well as the Unified Genotyper from GATK. For an interesting and informative workshop outlining the Broad best practices SNP calling pipeline check out these youtube videos (http://www.youtube.com/watch?v=1m0ZiEvzDKI&list=PLlMMtlgw6qNgNKNv5V9qmjAxbkHAZS1Mf). My results are in a series of blog posts and I hope you find them useful. Please let me know if you have any suggestions for SNP calling. We only want to do the alignments and SNP calling once for the entire set of samples, because it is going to take a long time!
GBS snp calling
One of the main ways we call SNPs in GBS data is using a maximum likelihood script derived from the original Hohenlohe stickleback RAD paper. It uses a likelihood function based on the number of reads for each base at a site to determine if its homozygous or heterozygous.
It’s been known for a while that it seemed to be biased against high coverage heterozygous sites. I took the script apart and realized that there was actually a serious problem with it. The likelihood function used factorials, and when there were over 175 reads at a site, perl broke down and started describing things as infinite. Then the script would call it as a homozygote by default.
Letting perl work with the huge numbers necessary makes it incredibly slow, so I put in a logic gate for high coverage sites. If there is over 150 reads at a site, and the second most numerous base makes up atleast 10% of the reads (i.e. at minimum 15), then it is heterozygous. Otherwise it is homozygous.
The rest of the script is the same. One could debate how best to call the high coverage scripts, but I suggest you use this script over the old one.
Filtering SNP tables
Previously I posted a pipeline for processing GBS data. At the end of the pipeline, there was a step where the loci were filtered for coverage and heterozygosity. How strict that filtering was could be changed, but it wasn’t obvious in the script and it was slow to rerun. I wrote a new script that takes a raw SNP table (with all sites, including ones with only one individual called) and calculates the stats for each loci. It is then simple and fast to use awk to filter this table for your individual needs. The only filtering the base script does is not print out sites that have four different bases called at the site.
Heterozygosity, read counts and GBS: PART 2
Subtitle: This time… its correlated.
Previously I showed that with the default ML snp calling on GBS data, heterozygosity was higher with high and low amounts of data. I then took my data, fed it through a snp-recaller which looks for sites that were called as homozygous but had at least 5 reads that matched another possible base at that position (i.e. a base that had been called there in another sample). I pulled all that data together, and put it into a single table with all samples where I filtered by:
Merge SNP calls (Greg B.)
Rose coded this up as a faster and efficient way to combine all the snp calls into one table. I’ve made a few modifications, hopefully its not broken. Updates are likely in the future.
SNP calling with ML (Greg B.)
Email for v7. Bug found that printed G’s as C’s and vise versa.
Call SNPs from sam files in a method similar to Hohenlohe et al 2010. Updated to v4 Feb 9. Previous version had a bug.
It is now fixed up for all of BWAs cigars flavors.
This only deals with reads that fit one of the following:
Full alignment (55M)
Soft clip at the start (10S45M)
Soft clip at end (45M10S)
One deletion (25M10D25M)
One insertion (20M10I20M)
This means it ignores reads with a cigar fields that have N, H, P, = or X and it ignores reads with a cigar more complicate then a single soft clip or a single indel. It also does not penalize reads adjacent to indels.
It ignores bases in soft clipped parts of reads
SNP table parsing (Greg B.)
Ask me for the most current version if you want to use any of these!
A few perl scripts that take a SNP table and do the following:
1. Remove unwanted samples and rename the samples
2. Remove sites that do not have enough data
3. Order the sites based on a map
Population Genomics! (Greg B.)
Ask me for the most current version of these if you want to use them.
Several people in the lab are now working with very similar data (in structure) and have similar questions (in technique). As I have discussed with several people it would be very useful if we could all build up and share the tools needed to do these analysis. Understandably, people may want to do things on their own or in a specific manner, but I think there are several advantages to building this up together. The main thing is that it will be more efficient in terms of personnel time, having each person re-invent the wheel does not make sense. I think the blog is a great place to set do this. Below we can make our wish list and link to posts for solutions as they become available. As always, the principles (1,2 and 3) covered here apply.