As some of you might know, I have been working for the last few months on optimizing a protocol for Illumina WGS libraries that will reduce our dependency on expensive kits without sacrificing quality. The ultimate goal would be to be able to use WGS libraries as a more expensive but hopefully more informative alternative genotyping tool to GBS. Getting to that point ideally requires to develop:
1) A cheaper alternative for library preparation (this post)
2) A reliable multiplexing system (this other post)
3) A way to shrink the sunflower genome before sequencing it (because, as you know, it’s rather huge) (yet another post)
The following protocol is for non-multiplexed libraries. The protocol for multiplexed ones is actually identical, you just need to change adapters and PCR primers – more about that in the multiplexing post.
If you are planning to pool libraries and deplete them of repetitive elements, read carefully all three posts before starting your libraries (mostly because you might need to use different adapters and PCR primers)
This protocol uses 1 μg of genomic DNA as starting material, and aims for 350bp-long fragments. Both can be changed (you can probably use much less DNA, if you are ok with increasing the number of PCR cycles in the enrichment step), and I added some notes about that at the end of the protocol. I’ll make a separate blog post about normalizing/quantifying your libraries by qPCR, without using the rather expensive kit form Kapa (here it is).
In its current form, this protocol allows to make high-quality libraries, from shearing to complete quality control, in one long-ish day (about 11 hours), without rushing it. This does not include normalization/quantification by qPCR, which would make it a rather long day (14-15 hours). All for less than 15 dollars/library, against 50+ dollars of the cheaper alternative that was used previously in the lab. I have to say, iI didn’t expect it to work quite so well.
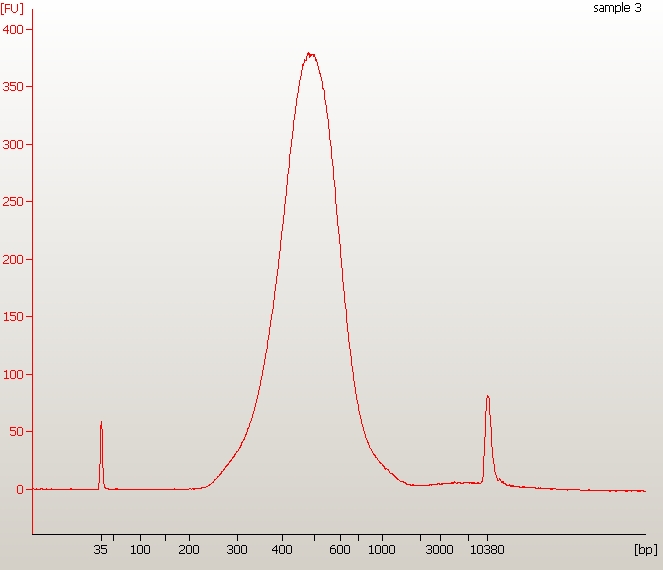
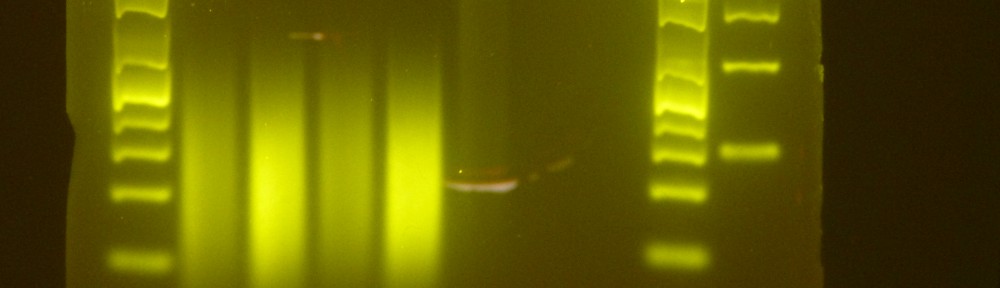
Example of home-brew Illumina library – All your libraries done with this protocol will look exactly like this one! (no warranty whatsoever)
I’ll soon organize a couple of boxes with all the required reagents (that is, if someone is interested in using this protocol). Please let me know if something is wrong or not clear with the protocol, or if you are interested in trying or discussing it.
Hi, I added some information on how long it takes to go through the whole protocol (not very long).
I uploaded an updated version of the protocol with a couple of corrections – mainly, I don’t use Phusion anymore, but the KAPA HiFi HotStart ReadyMix instead.
added v1.3 of the protocol, with some minor changes to the volumes of SPRI beads used in the purification steps
and v1.4 fixes a mistake in the the volume of water for the enrichment step using the Kapa HiFi polymerase.
and v1.5 adds a few note on how to get the amount of library you need for the DSN depletion treatment