Regardless of what method you use to make your Illumina libraries, if you added barcodes or indices you will need to normalize them before pooling (or otherwise have probably very uneven coverage). Or, you might want to know the exact molarity of your library before sending it for sequencing (although the need for that in our case is debatable, see later). The most accurate way to do both is probably by qPCR.
If you are not familiar with qPCR, following is a brief explanation of how it works. If you know all this stuff already, you can jump to the next chapter.
A brief introduction to qPCR
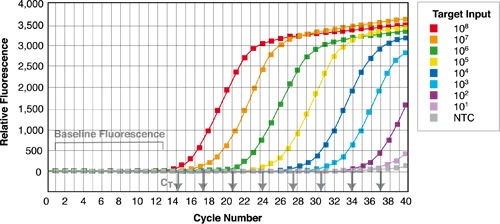
qPCR is just like a regular PCR, but occurs in the presence of a dye (generally SYBR green) which binds preferentially to double strand DNA as it gets synthesized. The level of fluorescence is recorder after each PCR cycle, and it will increase roughly two-fold each cycle (since after a successful PCR cycle you will have ideally double the amount of DNA for the sequence you are amplifying). The output you’ll get from a qPCR machine will look something like this (courtesy of Thermo Scientific):
For the first few cycles the levels of fluorescence will be below the detection threshold of the sensors. When you start to have enough DNA, the signal starts increasing exponentially, up to when it reaches saturation. As you can see in the example above, if you start with more of the sequence you want to amplify, the exponential phase will occur in earlier cycles. Using qPCR you can therefore compare the relative abundance of a certain sequence of DNA (the one you amplify with your specific primers) in different samples, or, if you have standards of known concentration, you can determine its absolute concentration.
QPCR is often used to quantify gene expression. Say for example you want to know the expression levels of the homeotic gene LFY in different growth stage of your plant; you take RNA from your samples, retro-transcribe it to get complementary DNA (cDNA), and then use it as a template for your qPCR (which in this case in generally called quantitative reverse transcription PCR, o qRT PCR). Using primers specific to the LFY sequence you’ll have an idea of how abundant the LFY transcript is in your different samples (you’ll need to run a separate qPCR on a housekeeping gene – whose expression supposedly is constant in different conditions – in order to properly normalize your results to the starting amount of RNA).
QPCR though can be used to quantify any other DNA sequence. For example, you can use primers specific for a nuclear gene and for a chloroplast gene on your genomic DNA to know how much chloroplast contamination you have in it. Or, you can use primers that recognize the adapters of your Illumina library to know how many amplifiable molecules you have in it (ehi, that’s exactly what we need!)
Normalizing your libraries by qPCR without buying a fancy kit
The most popular qPCR quantification kit for Illumina libraries in the Kapa Library Quantification Kit. It works very well and everybody is happy with it. It is also quite expensive, especially so if you don’t do a full plate of qPCRs every time. What is in the kit is a set of primers that recognize the extremities of Illumina adapters, some SYBR green PCR master mix, and a set of standards, which are basically sequential dilutions of a library of known molarity. The first two components are pretty standard and you can get them anywhere. The limiting ingredient in the kit is the standards, which are indeed very important if you want to know the exact absolute molarity of your libraries; quite intuitively, if your standards are not accurate, so will be your quantification.
As I mentioned before, I would argue though that, at least in our case, knowing the precise molarity of your final library is not paramount. We always outsource the sequencing itself, and any sequencing service will re-do some quality control and quantification of the libraries it receives. Still, they often ask you to send your libraries at a certain concentration, often express as molarity. This is because to optimize cluster density on a flowcell what matters is how many fragments you have, not the total amount of DNA you have in your library (which is the information you get out of a Qubit). Two libraries with the same concentration in mg/ml can have quite different molarity, if they have different fragment size. Just to normalize your libraries for pooling, you actually do not need to know their absolute molarity at all, just their relative ones. Knowing their absolute molarity does not affect the precision of your normalization, but still, after normalizing them you are going to pool them and send them for sequencing, so knowing the molarity will be useful at some point.
What I used instead of the Kapa kit were oligos I ordered from IDT and the Maxima SYBR Green qPCR Master Mix from Thermo Scientific, which works well and is reasonably priced. As standards I used a series of sequential dilutions of a library Dan Ebert made and quantified with the Kapa kit (average fragment size = 409 bp). At this stage your library will be quite concentrated, and beside you don’t want to waste too much of it as qPCR template. For my libraries I found that a 100.000 dilution gave you a final concentration that is around the middle of the range covered by the standards I used (which in turn had the same concentrations as the ones in the Kapa kit).
Setting up the qPCR reaction is actually easier than a normal PCR, since all you need is:
water 7 microl
SYBR green mix 10 microl
Illumina qPCR primer F 0.5 microl
Illumina qPCR primer R 0.5 microl
Diluted library 2 microl
Cycling conditions:
95°C 5 minutes
95°C 15 seconds
60°C 30 seconds for 40 cycles
72°C 30 seconds
data acquisition
72°C 5 minutes
Make three replicates for each library (and for each standard, if you use them). If you input the standard correctly, the qPCR machine will directly give you a concentration, normally in the tenths-of-pM range. Still, you have to consider that the average fragment size (which you get by running your libraries on a Bioanalyzer chip) is probably different between your libraries, and to the one of the standards. To calculate the actual molarity of your library therefore just use this formula:
Library = LibraryqPCR x (Fragment size of standards/Fragment size of your library) x 100.000
Where “Library” is the actual concentration of your library, “LibraryqPCR” is the concentration you obtained from your qPCR, and the rest is hopefully self-explanatory. As Greg O rightly pointed out, the number you get out of your qPCR is the molarity of the 100.000-fold dilution of your library. Therefore, you multiply it by 100.000 to get the actual concentration of your library.
So far the molarity values I got are almost identical to those that Ana got on the same libraries using the Kapa kit, so it works reasonably well. Similarly, when I pooled two libraries the sequencing output (as number of reads) was almost identical between them, so normalization with this method works well too (for less than a third of the most optimistic price estimate for the Kapa kit). If someone is interested in using the primers and standards, I’ll add them to the soon-to-be home-brew Illumina libraries reagents boxes.

Great post Marco. A couple questions:
-For the qPCR reaction, Illumina qPCR primer F is listed twice with different volumes. Is that supposed to be like that?
-What is the average fragment size of the library used for standards (assuming there is enough of the ebert library for others to use)?
Also, I think you should indicate that the 100,000 used in the library concentration equation is the dilution factor used.
Hi Greg, thanks for noticing the mistakes 🙂
– it is supposed to be like that only if you are lazy and copy the master mix formatting from another document, but forget to change the volumes. I fixed them. Also, the volumes are obviously not in milliliters, but wordpress doesn’t seem to like my “mu”, so i wrote microl for microliters.
– I added the average fragment size for Dan’s library (409 bp). I have about 1 ml for each dilution, which should be good for quantifying up to about 8500 libraries (so, you are welcome to use them :)).
and I clarified the 100.000 thing. Thanks again!