3rd generation sequencing technologies (PacBio, Oxford Nanopore) can produce reads that are several tens of Kb long. That is awesome, but it means that you also need to start from intact DNA molecules of at least that size. I thought the DNA extraction method I normally use for sunflower would be good enough, but that is not the case.
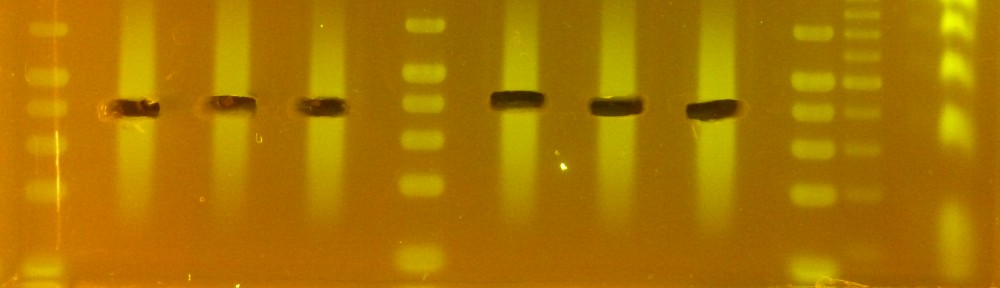
I have been using instead the protocol that was used to get DNA for sequencing the XRQ genome (Mayjonade et al. BioTechniques 2016), which is quick and works very well. I am posting here a commented version of the protocol. The main things I learned while testing this is that grinding the tissue with metal beads (as opposed to mortar and pestle) reduces DNA integrity, and that etiolated sunflower seedlings are not an easy tissue to get clean DNA from.