 The Parfrey lab investigates the diversity and distribution of microbial eukaryotes (protists)
The Parfrey lab investigates the diversity and distribution of microbial eukaryotes (protists)
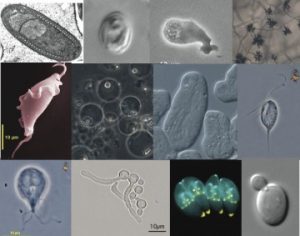
across environments. Eukaryotes are incredibly diverse and consist of many microbial lineages of amoebae, flagellates, ciliates and algae (plants, animals and fungi are also eukaryotes). We are broadly interested in understanding the factors the structure microbial communities, and in the interplay between microbial eukaryotes and bacteria. Much of our work focuses on host-associated communities in the mammalian gut and on coastal marine hosts. We use high-throughput sequencing to profile microbial communities combined with manipulative experiments to describe patterns of microbial diversity and uncover their underlying causes. Throughout, we use a phylogenetic framework to gain an evolutionary perspective on microbial community diversity.
Host associated microbial communities. All multicellular organisms host diverse communities of associated microbes. We investigate the diversity and assembly of microbial communities on a wide range of hosts, from coastal seaweeds and invertebrates to humans. High-throughput sequencing has revolutionized our understanding of host-associated microbes and demonstrated their widespread influence on host biology, ecology, and evolution.
Traditionally, eukaryotic microbes associated with animals, especially humans, were studied within the field of parasitology. As a result, microbial eukaryotes found in and on animals are considered parasites by default. However, only a subset are detrimental to their host—and are thus true parasites; many have neutral or beneficial effects on their host—they are commensals or mutualists. We refer to all host-associated eukaryotes as symbionts to acknowledge this diversity of host relationships.
Mammalian Microbiome
Our mammalian microbiome research strives to describe the normal community of eukaryotic microbes that are found in the mammalian gut. Microbes make up huge part of who we are as humans and influence many aspects of our lives ranging from disease, immune system development, nutrition and even behavior. Western life styles have greatly altered our microbial consortium and our health. These insights come almost exclusively from bacteria, but we have many reasons to suspect that eukaryotic microbes also play an important role at the community level. To begin to elucidate the role of eukaryotes within the microbiome community my lab is collaborating with other researchers to understand the normal consortia of eukaryotes within the human microbiome. With that knowledge we hope to characterize changes that have happened during westernization and in disease states. We are also assessing microbial eukaryotic communities in other vertebrate hosts to elucidate broader ecological and evolutionary patterns.

Coastal Host-Associated Microbiota
Microbes are found on all multicellular organisms and can profoundly influence host biology and broader ecosystem function. Our lab is part of several multi-disciplinary research projects characterizing microbial communities on coastal animals, plants, and seaweeds in British Columbia. These projects combine surveys that describe bacterial and eukaryotic microbial diversity patterns across host species with manipulative experiments to better understand microbial community assembly and the influence of microbes on host biology.
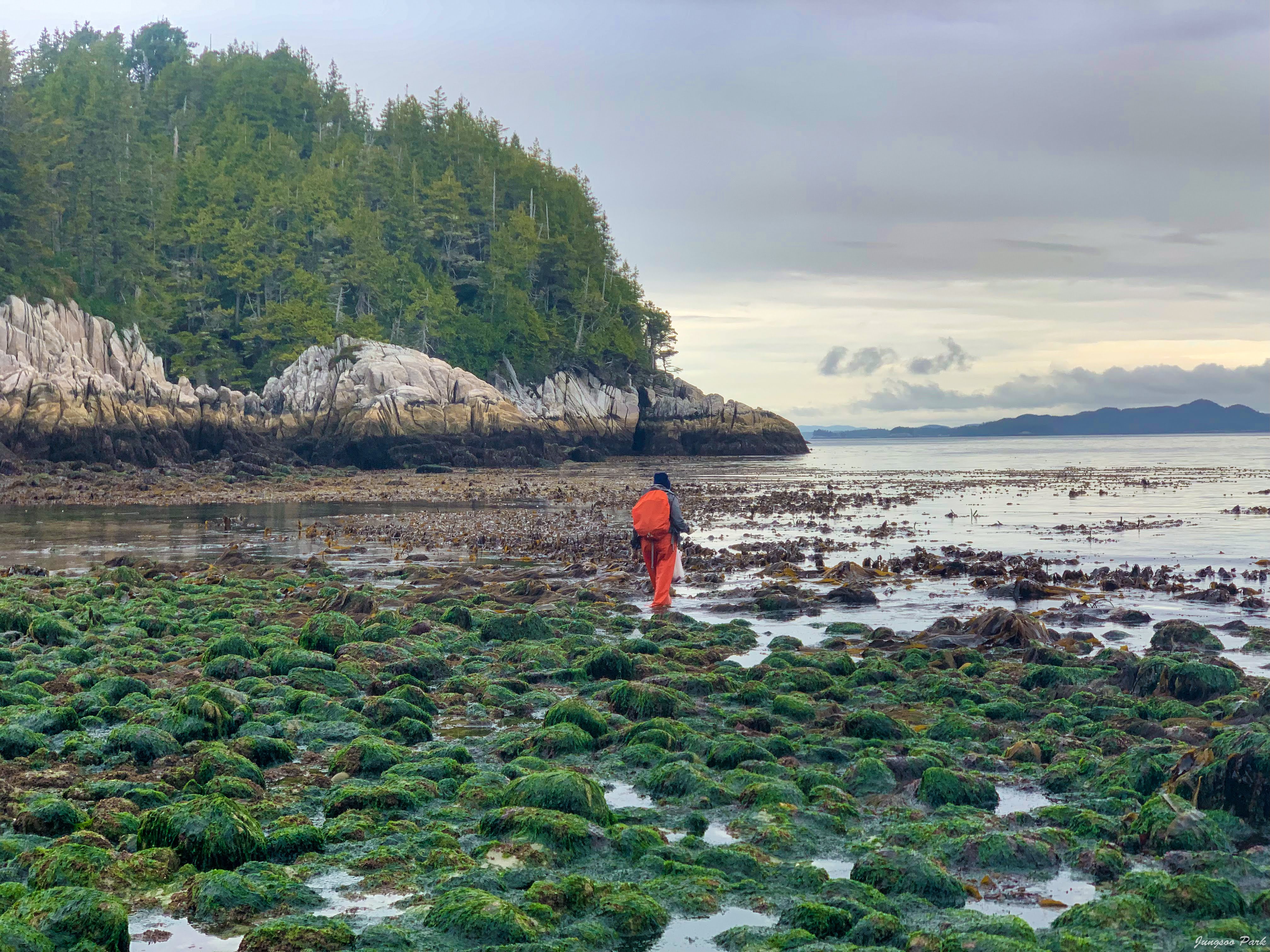
The Microbes to Macrophytes project seeks to quantify the interaction between ecologically important macrophytes (seagrass and seaweeds) and their associated microbes. This work takes place on and around Calvert Island on the central coast of British Columbia and is done in partnership with the Hakai Institute. Components include surveys across intertidal and subtidal seaweed species to understand community assembly and patterns of microbial diversity across a range of host species. We are also working with marine ecologists to understand the role of microbes within food webs and in carbon cycling within subtidal ecosystems. Our role is characterizing microbial diversity and function of communities found on the foundational species in seagrass meadows (Zostera marina) and kelp forests (Macrocystis pyrifera) in these subtidal ecosystems. Data from this project will be integrated into ongoing research to improve our understanding of marine community ecology and ecosystem function and provide insight into carbon cycling in intertidal and near-shore ecosystems.
Mapping epiphytic microbes on eelgrass leaves
A newly-formed collaboration between the Parfrey Lab and IMERSS
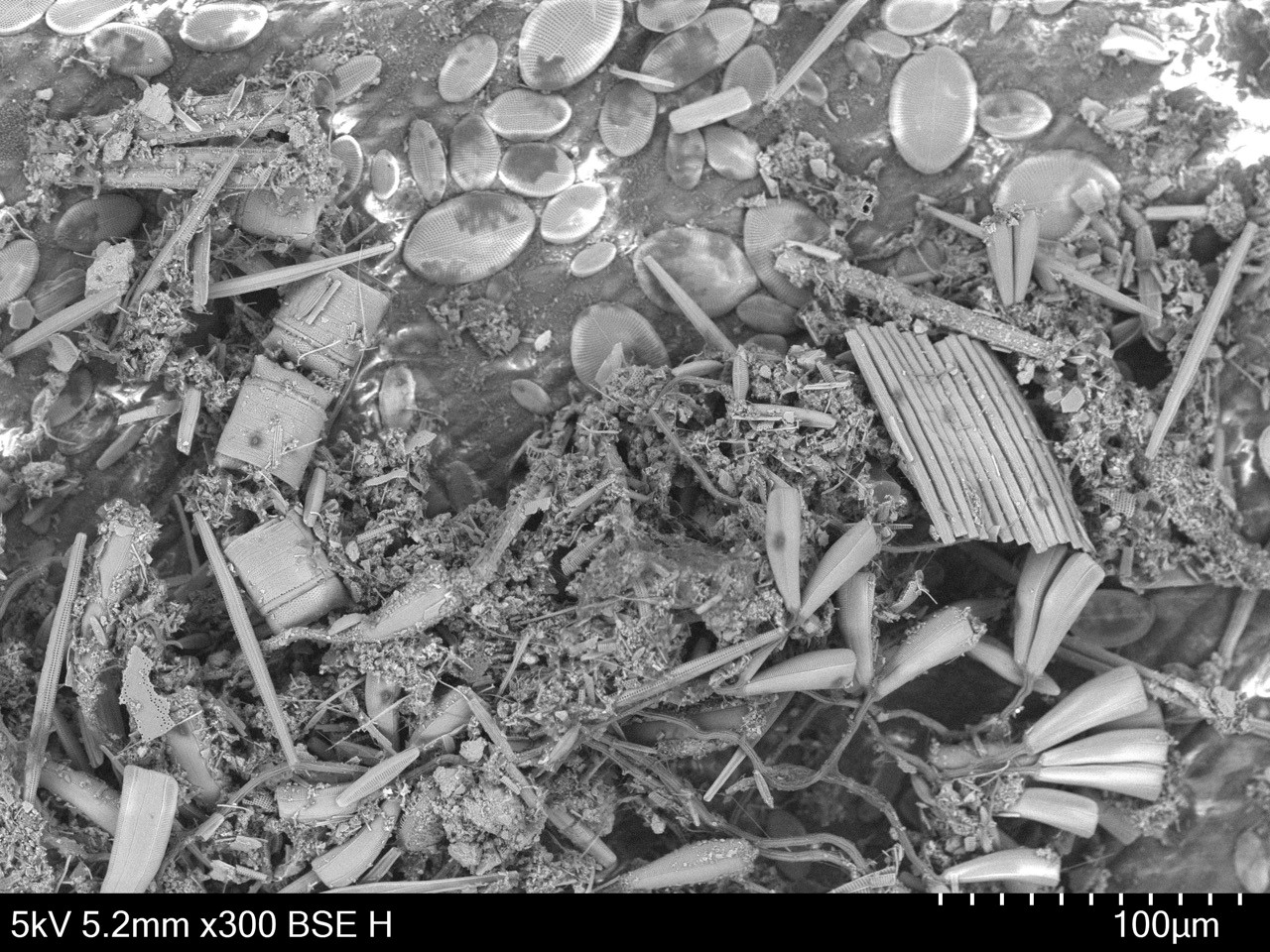

Eelgrass (Zostera marina) meadows are home to a diverse number of species, including fish, crabs, algae, and microorganisms. Eelgrass ecosystems contribute to many important coastal processes that humans rely on, such as carbon storage and coastal protection. Although eelgrass is well studied worldwide, there is little research that investigates the epiphytic microbes on leaf surfaces. Using scanning electron microscopy, we are mapping the surface of eelgrass leaves to visualize the spatial patterns and potential interactions of bacteria, fungi, and diatoms along the surface of the leaf. Additionally, we are cataloging bacterial, fungal, and diatom community diversity on eelgrass leaves and how diversity changes over time.
 Free-Living Eukaryotic and Bacterial Communities
Free-Living Eukaryotic and Bacterial Communities
We are investigating free-living protist and bacterial communities to better understand how selective factors influence free-living microbial communities. Jointly investigating protists and bacteria allows us to compare and contrast bacterial and eukaryotic diversity patterns. This approach also gives us a whole community perspective and can reveal interactions between the two domains. Two current projects investigate the influence of salinity and large protists on microbial community structure. The salinity project uses spatial and temporal surveys to characterize microbial changes across salinity gradients in the Fraser River Estuary and the Straight of Georgia around Vancouver, while we use manipulative experiments in the lab to test the effects of large protists.
 Improving Infrastructure for Analyses of Microbial Eukaryotes
Improving Infrastructure for Analyses of Microbial Eukaryotes
High throughput sequencing has dramatically expanded our knowledge of protist biodiversity and their distribution across environments. However, these studies rely on reference databases to annotate sequences and make sense of the vast amounts of data produced. The quality of eukaryotic reference databases (primarily SILVA and PR2) has improved in recent years, but still struggles to keep pace with the rapid influx of data, new lineages, and changing eukaryotic classification. With collaborators, we are leading a community-wide effort (EukRef) that brings together taxonomists and experts in lineages from across the eukaryotic tree of life to curate high quality and phylogenetically informed reference databases. The Parfrey lab is also working to improve tools for eukaryotic community analysis.
*Photo used under creative commons license from Encyclopedia of Life and Wiki Commons