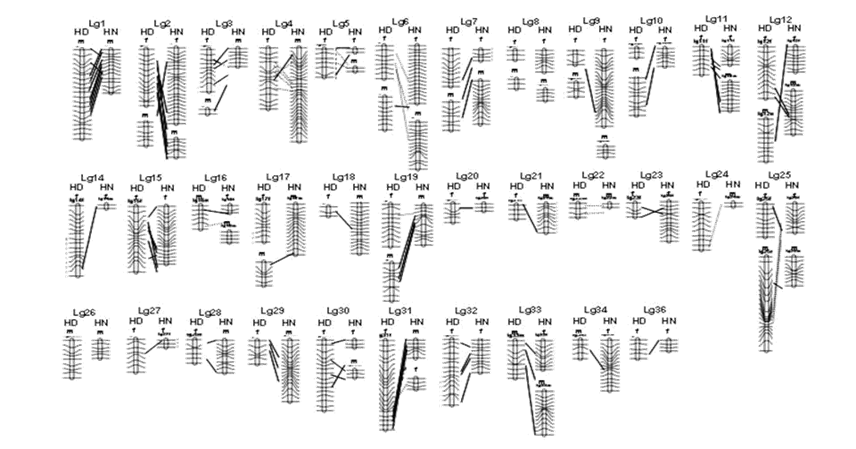Sean M. Rogers
|
|
Research
My research has therefore
focused on bridging the gap between ecology and genetics towards
understanding how organisms adapt to new and changing environments. I use
fish species to answer these questions for primarily two reasons. First, many
of the fish populations in
|
|
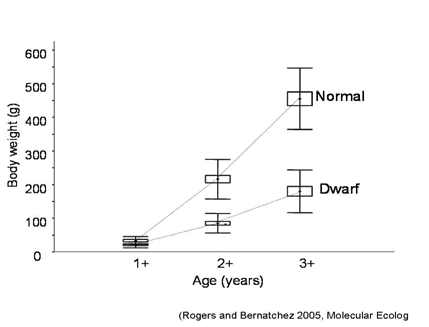
Dwarf
and I worked
with the lake whitefish (Coregonus clupeaformis, Mitchill) for my PhD thesis. These amazing fish are in the
same family as trout and salmon and inhabit coldwater lakes all over the
Northern hemisphere. In several lakes across The objective of my PhD thesis
was to determine the genetic basis of the differences between dwarf and
normal lake whitefish. This would help us understand the genetic changes that
occur during ecological divergence and adaptation to new environments.
Specifically, one of my objectives was to localize regions of the whitefish
genome that were associated with adaptation and reproductive isolation
between dwarf and normal populations. This would give us clues as to what
evolutionary steps have led to the inability (or reduced ability) of these
two populations to interbreed freely. To achieve this objective, I first made
two hybrid backcrosses between dwarf and normal whitefish. Progeny from these
crosses were sampled for their DNA and individually tagged (with PIT
tags). This allowed me to study the inheritance of several hundred genetic
markers in the progeny of both families and construct a genetic map. The
genetic map provided the necessary template to localize genomic regions
associated with several behavioural, physiological,
and morphological traits that differentiated dwarf and normal lake whitefish.
The genetic architecture of
population divergence: Linkage maps of the dwarf and I measured many traits that I
had identified as phenotype-environment associations and were thus associated
with adaptation and reproductive isolation. This involved performing
experiments on all of these fish throughout their life history, included
swimming behaviour video experiments, intensive
growth measurements from the juvenile stage to sexual maturity (over three
years), and also measurements of size, shape, and the sex of each individual.
At the end of the experiments, fish were frozen at -80 degrees Celsius to
preserve their RNA (for subsequent gene expression profiling using microarrays which is currently being done in
collaboration with Dr. Nicolas Derome and Dr. Andrew Whiteley).
Using quantitative trait locus
(QTL) mapping, I tested for associations between these traits and the genes
the progeny had inherited using the genetic map. I found several regions of
the genome that were linked in association with adaptive traits. More
importantly, there was evidence that natural selection had influenced these
regions among independently evolving species pairs inhabiting distinct lakes.
The loci exhibiting a signature of selection were more likely to be
associated with adaptive QTL than with other regions of the genome. This was
important because it offered evidence that natural selection at these key
genomic regions has been maintaining the differences between dwarf and normal
lake whitefish and contributing to the rapid evolution of these species
pairs. Much of this work was pioneered by Dr.
Louis Bernatchez who continues to work on nearly all aspects of
whitefish evolution, especially in genomics.
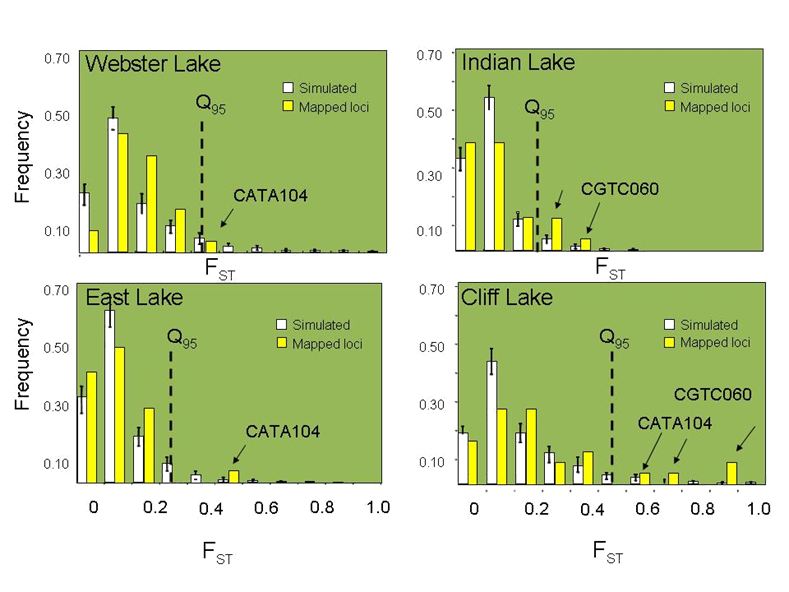
Genome scans of differential
gene exchange for growth associated mapped QTL (yellow bars) among four
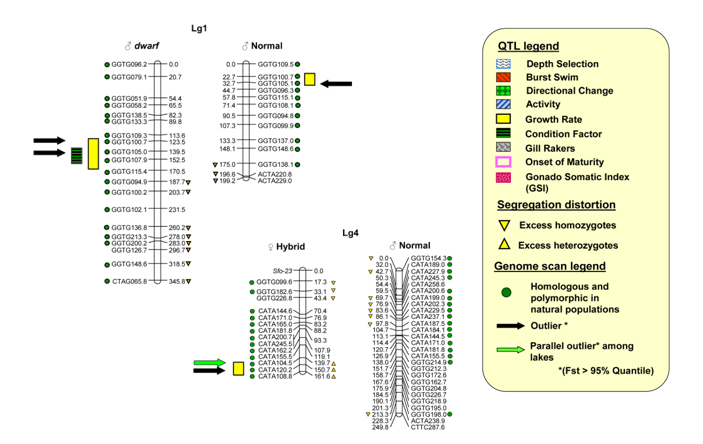
natural sympatric pairs of dwarf and normal lake whitefish (a) Genetic maps of whitefish
linkage groups 1 and 4 showing the locations of adaptive QTL overlapping with
outlier loci that exhibit a signature of selection among natural sympatric
pairs of lake whitefish (Rogers and Bernatchez, Molecular Biology and
Evolution 2007) Current Research:
Photo: Kerry
Marchinko With the collaboration of the Stanford Genome
Evolution Center, I am conducting experiments that apply the
theory for the genetics of adaptation to four wild populations. This
experiment involves repeatedly crossing the same marine ancestor to fish from
four recently colonized freshwater lakes that differ in age and degree of
phenotypic divergence. Genetic and molecular tools in these crosses will be
used to test predictions about the genetics of parallel adaptive trait
divergence. Comparing the genetic basis of adaptations in the older, more
divergent lakes with the QTL underlying younger, more recent colonizations will allow us to test specific predictions
about the genetic changes that have occurred during the course of freshwater
evolution. I am also collaborating with Rowan
Barrett, a PhD student in Dolph Schluter’s
lab. We are experimentally testing the genetics of adaptation from standing
variation by colonizing freshwater experimental ponds with natural marine
fish that have been genetically confirmed to have one copy of an ancient
allele at the key candidate gene (Eda)
underlying freshwater armour evolution in
sticklebacks. This ancient allele occurs at a very low frequency in saltwater
(only 1/1000 marine sticklebacks are homozygous for the low allele), but it
is fixed in most freshwater stickleback populations (Colosimo
et al. 2005). Because we have colonized our ponds with marine sticklebacks
that are heterozygous for Eda, we are able
to predict the starting frequency of the genotypes in the next generation in
the absence of selection.
Evolution ponds (photo by
Rowan Barrett) |
|
|
|
|
|
|