plot of chunk unnamed-chunk-2
| make.bisse.td {diversitree} | R Documentation |
Create a likelihood function for a BiSSE model where different chunks of time have different parameters. This code is experimental!
make.bisse.td(tree, states, n.epoch, unresolved=NULL, sampling.f=NULL,
nt.extra=10, strict=TRUE, control=list())
make.bisse.t(tree, states, functions, unresolved=NULL, sampling.f=NULL,
nt.extra=10, strict=TRUE, control=list())
tree |
An ultrametric bifurcating phylogenetic tree, in
|
states |
A vector of character states, each of which must be 0 or
1, or |
n.epoch |
Number of epochs. 1 corresponds to plain BiSSE, so this will generally be an integer at least 2. |
functions |
A named list of functions of time. See details. |
unresolved |
Unresolved clade information: see
|
sampling.f |
Vector of length 2 with the estimated proportion of
extant species in state 0 and 1 that are included in the phylogeny.
See |
nt.extra |
The number of species modelled in unresolved clades (this is in addition to the largest observed clade). |
strict |
The |
control |
List of control parameters for the ODE solver. See
details in |
This builds a BiSSE likelihood function where different regions of time (epochs) have different parameter sets. By default, all parameters are free to vary between epochs, so some constraining will probably be required to get reasonable answers.
For n epochs, there are n-1 time points; the first
n-1 elements of the likelihood's parameter vector are these
points. These are measured from the present at time zero, with time
increasing towards the base of the tree. The rest of the parameter
vector are BiSSE parameters; the elements n:(n+6) are for the
first epoch (closest to the present), elements (n+7):(n+13) are
for the second epoch, and so on.
For make.bisse.t, the funtions is a list of functions
of time. For example, to have speciation rates be linear functions of
time, while the extinction and character change rates be constant with
respect to time, one can do
functions=rep(list(linear.t, constant.t), c(2, 4))The functions here must have
t as their first argument,
interpreted as time back from the present. See
make.pars.t for more information, and for some
potentially useful time functions.
Richard G. FitzJohn
set.seed(4)
pars <- c(0.1, 0.2, 0.03, 0.03, 0.01, 0.01)
phy <- tree.bisse(pars, max.t = 30, x0 = 0)Suppose we want to see if diversification is different in the most recent 3 time units, compared with the rest of the tree (yes, this is a totally contrived example!):
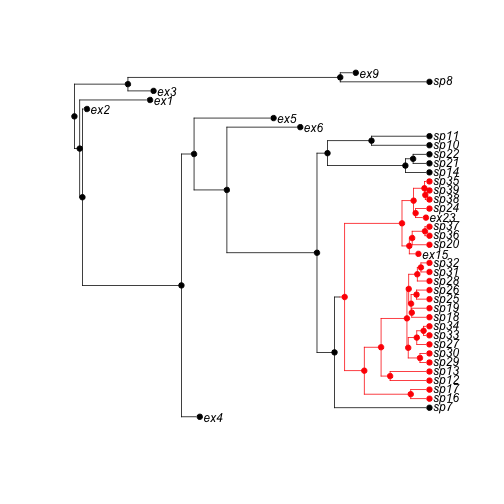
plot(phy)
axisPhylo()
abline(v = max(branching.times(phy)) - 3, col = "red", lty = 3)
plot of chunk unnamed-chunk-2
For comparison, make a plain BiSSE likelihood function
lik.b <- make.bisse(phy, phy$tip.state)Create the time-dependent likelihood function. The final argument here is the number of 'epochs' that are allowed. Two epochs is one switch point.
lik.t <- make.bisse.td(phy, phy$tip.state, 2)The switch point is the first argument. The remaining 12 parameters are the BiSSE parameters, with the first 6 being the most recent epoch.
argnames(lik.t)## [1] "t.1" "lambda0.1" "lambda1.1" "mu0.1" "mu1.1"
## [6] "q01.1" "q10.1" "lambda0.2" "lambda1.2" "mu0.2"
## [11] "mu1.2" "q01.2" "q10.2"
pars.t <- c(3, pars, pars)
names(pars.t) <- argnames(lik.t)Calculations are identical to a reasonable tolerance:
lik.b(pars) - lik.t(pars.t)## [1] -8.144e-07It will often be useful to constrain the time as a fixed quantity.
lik.t2 <- constrain(lik.t, t.1 ~ 3)Parameter estimation under maximum likelihood. This is marked "don't run" because the time-dependent fit takes a few minutes.
[This section is not run by default] Fit the BiSSE ML model
fit.b <- find.mle(lik.b, pars)And fit the BiSSE/td model
fit.t <- find.mle(lik.t2, pars.t[argnames(lik.t2)], control = list(maxit = 20000))Compare these two fits with a likelihood ratio test (lik.t2 is nested within lik.b)
anova(fit.b, td = fit.t)[Ends not run section]
The time varying model (bisse.t) is more general, but substantially slower. Here, I will show that the two functions are equivalent for step function models. We'll constrain all the non-lambda parameters to be the same over a time-switch at t=5. This leaves 8 parameters.
lik.td <- make.bisse.td(phy, phy$tip.state, 2)
lik.td2 <- constrain(lik.td, t.1 ~ 5, mu0.2 ~ mu0.1, mu1.2 ~ mu1.1,
q01.2 ~ q01.1, q10.2 ~ q10.1)
lik.t <- make.bisse.t(phy, phy$tip.state, rep(list(stepf.t, constant.t),
c(2, 4)))
lik.t2 <- constrain(lik.t, lambda0.tc ~ 5, lambda1.tc ~ 5)Note that the argument names for these functions are different from one another. This reflects different ways that the functions will tend to be used, but is potentially confusing here.
argnames(lik.td2)## [1] "lambda0.1" "lambda1.1" "mu0.1" "mu1.1" "q01.1" "q10.1"
## [7] "lambda0.2" "lambda1.2"argnames(lik.t2)## [1] "lambda0.y0" "lambda0.y1" "lambda1.y0" "lambda1.y1" "mu0"
## [6] "mu1" "q01" "q10" First, evaluate the functions with no time effect and check that they are the same as the base BiSSE model
p.td <- c(pars, pars[1:2])
p.t <- pars[c(1, 1, 2, 2, 3:6)]All agree:
lik.b(pars) # -159.7128## [1] -159.7lik.td2(p.td) # -159.7128## [1] -159.7lik.t2(p.t) # -159.7128## [1] -159.7In fact, the time-varying BiSSE will tend to be identical to plain BiSSE where the functions to not change:
lik.b(pars) - lik.t2(p.t)## [1] 0Slight numerical differences are typical for the time-chunk BiSSE, because it forces the integration to be carried out more carefully around the switch point.
lik.b(pars) - lik.td2(p.td)## [1] -1.2e-06Next, evaluate the functions with a time effect (5 time units ago, speciation rates were twice the contemporary rate)
p.td2 <- c(pars, pars[1:2] * 2)
p.t2 <- c(pars[1], pars[1] * 2, pars[2], pars[2] * 2, pars[3:6])Huge drop in the likelihood (from -159.7128 to -172.7874)
lik.b(pars)## [1] -159.7lik.td2(p.td2)## [1] -172.8lik.t2(p.t2)## [1] -172.8The small difference remains between the two approaches, but they are basically the same.
lik.td2(p.td2) - lik.t2(p.t2)## [1] 3.553e-06There is a time cost to both time-dependent methods, but it is more heavily paid for the time-varying case:
system.time(lik.b(pars))## user system elapsed
## 0.007 0.000 0.007 system.time(lik.td2(p.td)) # 1.9x slower than plain BiSSE## user system elapsed
## 0.013 0.000 0.013 system.time(lik.t2(p.t)) # 10x slower than plain BiSSE## user system elapsed
## 0.093 0.002 0.096